
I work on an interdisciplinary project on light regulation of gene expression and inter-tissue signalling in A. thaliana. In this project I combine molecular and computational biology approaches to answer how light is perceived in different tissues and how the downstream signalling is affected.
ABSTRACT
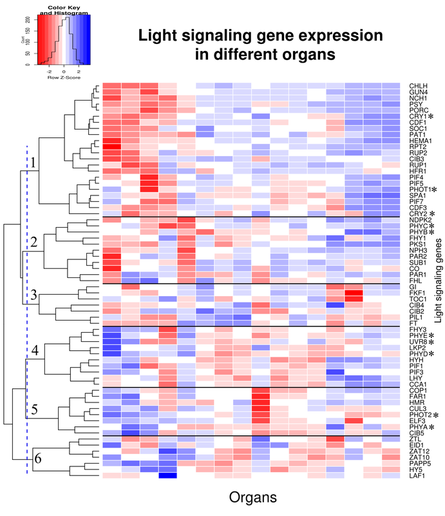
In the current research, whole seedling or leaf extracts are routinely investigated in microarray or RNAseq experiments to provide insight into changing levels of transcription of multiple genes of interest. However, in 2014 it was shown that the whole leaf extract is composed in majority of the mesophyll tissue (~80%) and gene expression resembles that of mesophyll (Endo et al., 2014). In this situation, gene expression from vascular tissue is largely underrepresented and information about tissue-specific expression is lost. Many examples of tissue specific and inter-tissue signalling in A. thaliana pose a question of whether the light signalling components are present in all cell types and if they are present in the same quantities. Although phytochromes and other receptors have been shown to have stable levels of expression among different organs (Sharrock and Clack, 2002), it is possible that dissecting their expression by tissue rather than organ will highlight differences in levels of light signalling components. In my project I aim to investigate the differences in light signalling pathways in various tissues and determine how these differences affect light perception and downstream effects. Through analysis of existing microarray and RNAseq datasets I will investigate whether components of light signalling network show variable expression across different organs and developmental stages and hypothesise that any variations between organs could suggest even more pronounced differences between tissues.
